4.7. Least squares model analysis¶
Once we have fitted the
Analysis of variance: breaking down the data’s variability into components
Confidence intervals for the model coefficients,
Prediction error estimates for the
We will also take a look at the interpretation of the software output.
In order to perform the second part we need to make a few assumptions about the data, and if the data follow those assumptions, then we can derive confidence intervals for the model parameters.
4.7.1. The variance breakdown¶
Recall that variability is what makes our data interesting. Without variance (i.e. just flat lines) we would have nothing to do. The analysis of variance is just a tool to show how much variability in the
Doing nothing (no model: this implies
) The model (
) How much variance is left over in the errors,
These 3 components must add up to the total variance we started with. By definition, the variance is computed about a mean, so the variance of no model (i.e. the “doing nothing” case) is zero. So the total variance in vector
Using the accompanying figure, we see that geometrically, at any fixed value of

The total sum of squares (TSS) is the total variance in the vector of
It is convenient to write these sums of squares (variances) in table form, called an Analysis of Variance (ANOVA) table:
Type of variance
Distance
Degrees of freedom
SSQ
Mean square
Regression
( in the examples so far) RegSS
Error
RSS
Total
TSS
4.7.1.1. Interpreting the standard error¶
The term
Example: Assume we have a model for predicting batch yield in kilograms from
Answer: Recall if the assumption of normally distributed errors is correct, then this value of 3.4 kg indicates that about two thirds of the yield predictions will lie within
4.7.1.2. Exercise¶
For two extreme cases:
Do the following in the space below:
draw a generic plot
create an ANOVA table with fake values
write down the value of the ratio
interpret what this ratio means:
From this exercise we learn that:
The null model (
Models where the fit is perfect have a ratio
4.7.1.3. Derivation of
As introduced by example in the previous part,
From the above ratios it is straightforward to see that if
The nomenclature
and can range in value from
The
These two common examples illustrate the abuse. You likely have said or heard something like this before:
“the
value is really high, 90%, so this is a good model”. “Wow, that’s a really low
, this model can’t be right - it’s no good”.
How good, or how suitable a model is for a particular purpose is almost never related to the
your engineering judgment: does the interpretation of model parameters make sense?
use testing data to verify the model’s predictive performance,
using cross-validation tools (we will see this topic later on) to see how well the model performs on new, unseen and unused testing data.
We will see later on that
where
4.7.2. Confidence intervals for the model coefficients
Note
A good reference for this section is the book by Fox (Chapter 6), and the book by Draper and Smith.
Up to this point we have made no assumptions about the data. In fact we can calculate the model estimates,
Recall the
We first take a look at some assumptions in least squares modelling, then return to deriving the confidence interval.
4.7.2.1. Assumptions required for analysis of the least squares model¶
Recall that the population (true) model is
Furthermore, our derivation for the confidence intervals of
Linearity of the model, and that the values of
In an engineering situation this would mean that your
The variance of
The variability of
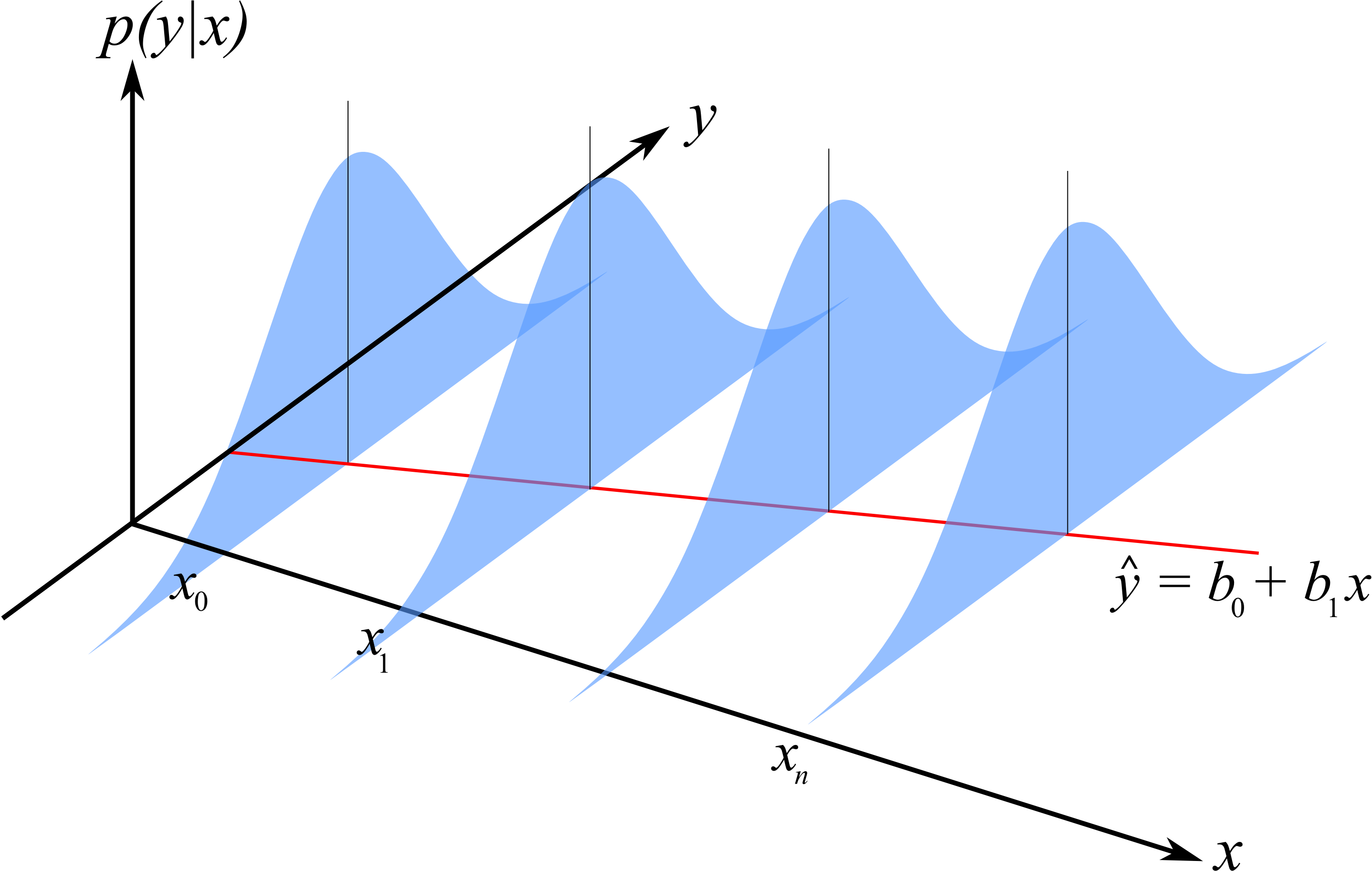
Illustration of the constant error variance assumption and the normally distributed error assumption.
The errors are normally distributed:
Each error is independent of the other. This assumption is often violated in cases where the observations are taken in time order on slow moving processes (e.g. if you have a positive error now, your next sample is also likely to have a positive error). We will have more to say about this later when we check for independence with an autocorrelation test.
In addition to the fact that the
When the
All
Note
Derivation of the model’s coefficients do not require these assumptions, only the derivation of the coefficient’s confidence intervals require this.
Also, if we want to interpret the model’s
4.7.2.2. Confidence intervals for
Recall from our discussions on confidence intervals that we need to know the mean and variance of the population from which
Once we know those parameters, we can create a
Start from the equations that define
That last form of expressing
So we can write:
where
Questions:
So now apart from the numerator term, how could you decrease the error in your model’s
Use samples that are far from the mean of the
Use more samples.
What do we use for the numerator term
This term represents the variance of the
Now for the variance of
Summary of important equations
For convenience we will define some short-hand notation, which is common in least squares:
You will see that
Now it is straight forward to construct confidence intervals for the least squares model parameters. You will also realize that we have to use the
Example
Returning back to our ongoing example, we can calculate the confidence interval for
Use that qt(0.975, df=(N-2)). There are
First calculate the
The 95% confidence interval for
The confidence interval for
The plot shows the effect of varying the slope parameter,
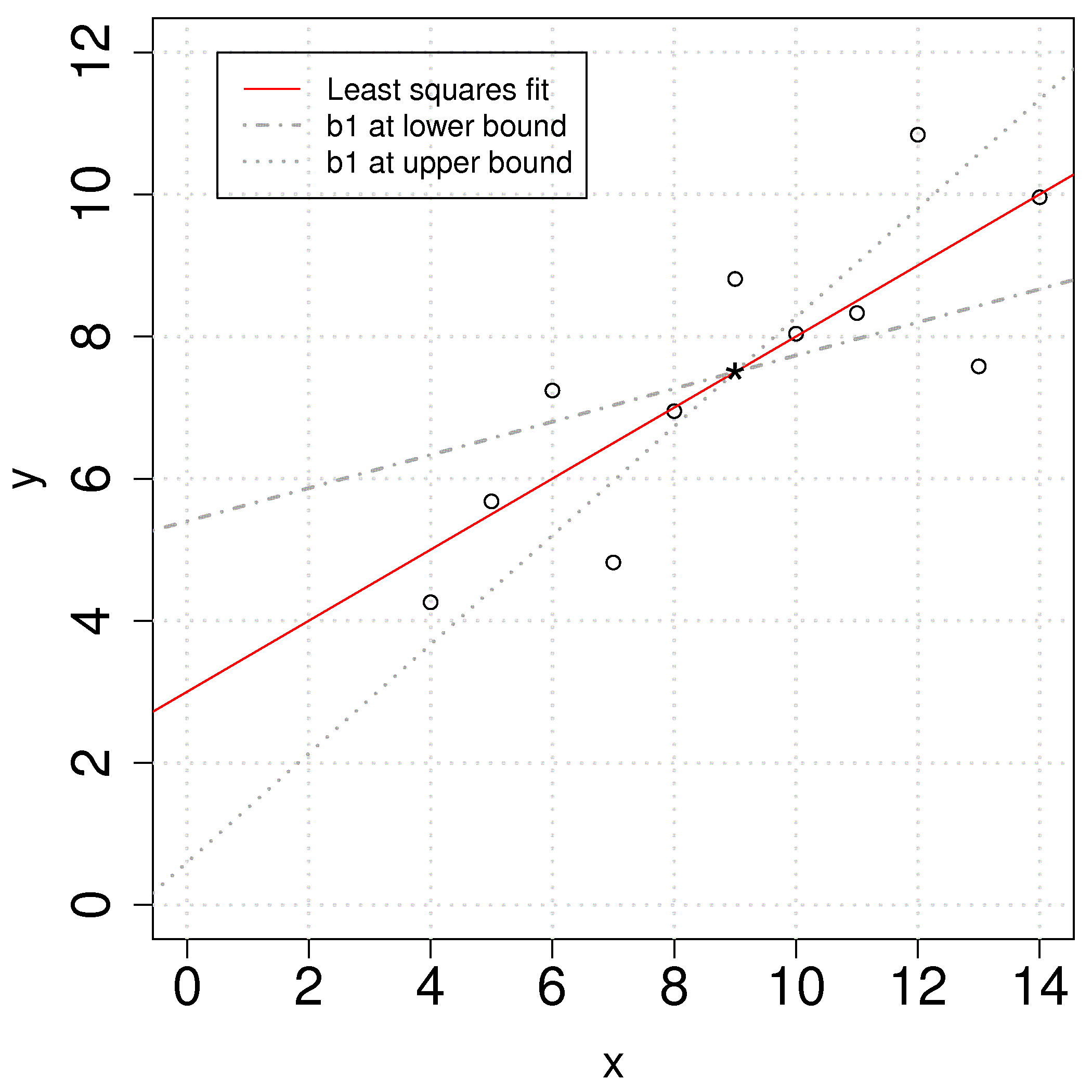
In many cases the confidence interval for the intercept is not of any value because the data for
4.7.3. Prediction error estimates for the y-variable¶
Apart from understanding the error in the model’s coefficient, we also would like an estimate of the error when predicting
4.7.3.1. A naive first attempt¶
We might expect the error is related to the average size of the residuals. After all, our assumptions we made earlier showed the standard error of the residuals was the standard error of the
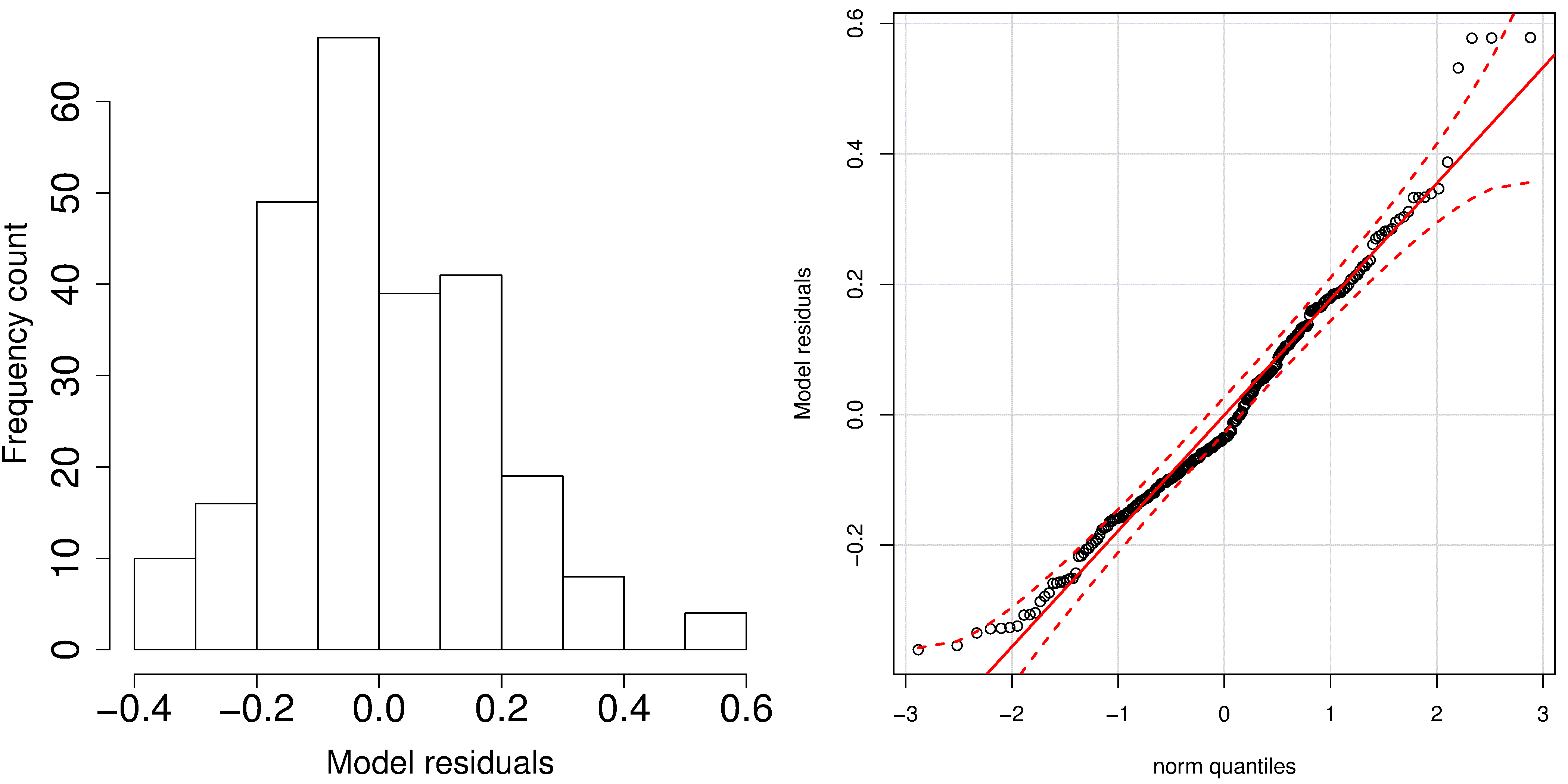
A typical histogram of the residuals looks as shown here: it is always centered around zero, and appears to be normally distributed. So we could expect to write our prediction error as
But there is something wrong with that error estimate. It says that our prediction error is constant at any value of
This estimate is however a reasonable guess for the prediction interval when you only know the model’s
4.7.3.2. A better attempt to construct prediction intervals for the least squares model¶
Note
A good reference for this section is Draper and Smith, Applied Regression Analysis, page 79.
The derivation for the prediction interval is similar to that for
You may read the reference texts for the interesting derivation of this variance. However, this is only the variance of the average predicted value of
We may construct a prediction interval in the standard manner, assuming that
This is a prediction interval for a new prediction,
4.7.3.3. Implications of the prediction error of a new
Let’s understand the interpretation of
Now let’s say our
The LB =
The UB =
Now move left and right, away from
The confidence intervals have a quadratic shape due to the square term under the square root. The smallest prediction error will always occur at the center of the model, and expands progressively wider as one moves away from the model center. This is illustrated in the figure and makes intuitive sense as well.
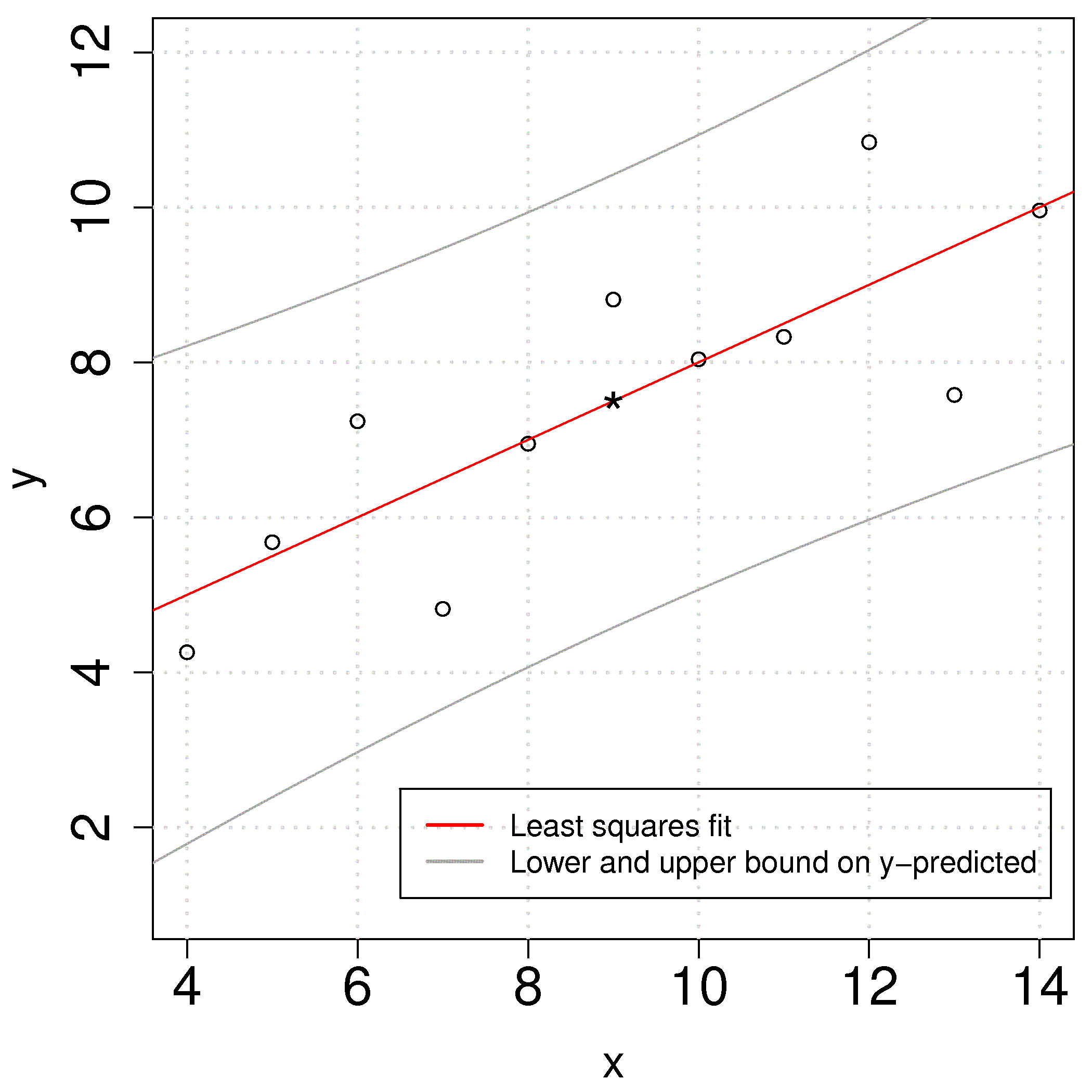
4.7.4. Interpretation of software output¶
To complete this section we show how to interpret the output from computer software packages. Most packages have very standardized output, and you should make sure that whatever package you use, that you can interpret the estimates of the parameters, their confidence intervals and get a feeling for the model’s performance.
The following output is obtained in R for the example we have been using in this section. The Python version follows below.
and produces this output:
Call:
lm(formula = y ~ x)
Residuals:
Min 1Q Median 3Q Max
-1.92127 -0.45577 -0.04136 0.70941 1.83882
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 3.0001 1.1247 2.667 0.02573 *
x 0.5001 0.1179 4.241 0.00217 **
---
Signif. codes: 0 `***' 0.001 `**' 0.01 `*' 0.05 `.' 0.1 ` ' 1
Residual standard error: 1.237 on 9 degrees of freedom
Multiple R-squared: 0.6665, Adjusted R-squared: 0.6295
F-statistic: 17.99 on 1 and 9 DF, p-value: 0.002170
Make sure you can calculate the following values using the equations developed so far, based on the above software output:
The intercept term
= 3.0001. The slope term
= 0.5001. The standard error of the model,
= 1.237, using degrees of freedom. Using the standard error, calculate the standard error for the intercept =
. Using the standard error, calculate the standard error for the slope =
. The
-value for the term is 2.667 (R calls this the t valuein the printout, but in our notes we have called this; the value that we compare to the -statistic and used to create the confidence interval). The
-value for the term is 4.241 (see the above comment again). The two probability values,
Pr(>|t|), forand should be familiar to you; they are the probability with which we expect to find a value of greater than the calculated -value (called t valuein the output above). The smaller the number, the more confident we can be the confidence interval contains the parameter estimate.You can construct the confidence interval for
or by using their reported standard errors and multiplying by the corresponding -value. For example, if you want 99% confidence limits, then look up the 99% values for the -distribution using degrees of freedom, in this case it would be qt((1-0.99)/2, df=9), which is. So the 99% confidence limits for the slope coefficient would be . The
value. Be able to calculate the residuals:
. We expect the median of the residuals to be around 0, and the rest of the summary of the residuals gives a feeling for how far the residuals range about zero.
Using Python, you can run the following code:
which produces the following output:
OLS Regression Results
==============================================================================
Dep. Variable: y R-squared: 0.667
Model: OLS Adj. R-squared: 0.629
Method: Least Squares F-statistic: 17.99
Date: Tue, 01 Jan 2019 Prob (F-statistic): 0.00217
Time: 00:00:00 Log-Likelihood: -16.841
No. Observations: 11 AIC: 37.68
Df Residuals: 9 BIC: 38.48
Df Model: 1
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
const 3.0001 1.125 2.667 0.026 0.456 5.544
x1 0.5001 0.118 4.241 0.002 0.233 0.767
==============================================================================
Omnibus: 0.082 Durbin-Watson: 3.212
Prob(Omnibus): 0.960 Jarque-Bera (JB): 0.289
Skew: -0.122 Prob(JB): 0.865
Kurtosis: 2.244 Cond. No. 29.1
==============================================================================
Standard error = 1.2366033227263207
As for the R code, we can see at a glance:
The intercept term
= 3.0001. The slope term
= 0.5001. The standard error of the model,
= 1.237, using degrees of freedom. The summary output table does not show the standard error, but you can get it from np.sqrt(results.scale), whereresultsis the Python object from fitting the linear model.Using the standard error, calculate the standard error for the intercept =
, which is reported directly in the table. Using the standard error, calculate the standard error for the slope =
, which is reported directly in the table. The
-value for the term is 2.667 (Python calls this the t-value in the printout, but in our notes we have called this; the value that we compare to the -statistic and used to create the confidence interval). The
-value for the term is 4.241 (see the above comment again). The two probability values,
P>|t|, forand should be familiar to you; they are the probability with which we expect to find a value of greater than the calculated -value (called t valuein the output above). The smaller the number, the more confident we can be the confidence interval contains the parameter estimate.You can construct the confidence interval for
or by using their reported standard errors and multiplying by the corresponding -value. For example, if you want 99% confidence limits, then look up the 99% values for the -distribution using degrees of freedom, in this case it would be from scipy.stats import t; t.ppf(1-(1-0.99)/2, df=9), which is. So the 99% confidence limits for the slope coefficient would be . However, the table output gives you the 95% confidence interval. Under the column 0.025and0.975(leaving 2.5% in the lower and upper tail respectively). For the slope coefficient, for example, this interval is [0.233; 0.767]. If you desire, for example, the 99% confidence interval, you can adjust the code:print(results.summary(alpha=1-0.99))The
value. Be able to calculate the residuals:
.